Twisty “Knot” DNA Discovered in Living Cells for the First Time
This unusual DNA is typically present in the wild and has *knot* been seen in living cells hitherto. The traditional double helix DNA occurring in living cells has now got an upgrade- a twisted knot upgrade.
Human genome function is underpinned by the primary storage of genetic information in canonical B-form DNA, with a second layer of DNA structure providing regulatory control. I-motif structures are thought to form in cytosine-rich regions of the genome and to have regulatory functions; however, in vivo evidence for the existence of such structures has so far remained elusive.
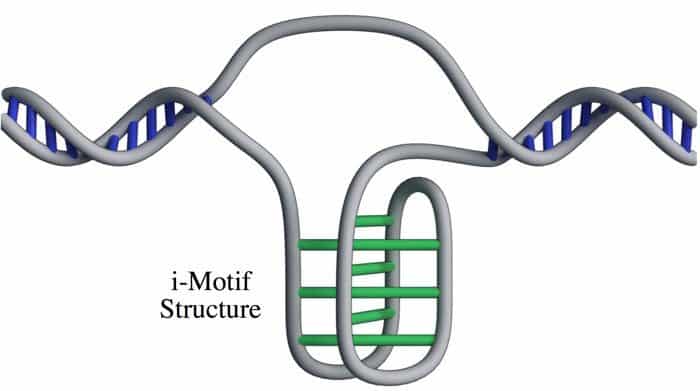
Scientists have now discovered a new structure inside human cells: a never-before-seen twisted “knot” of DNA. Dubbed the “i-motif,” this four-stranded knot looks totally different from the iconic double helix. This discovery in living cells confirms our complex genetic code is crafted with more intricate symmetry than just the double helix structure everybody associates with DNA – and the forms these molecular variants take affect how our biology functions.
The twisted knot structure only occurs in a relatively small region of a genome, like a knot in the helical double strands of DNA. In the twisted knot structure, Cs bind to Cs instead of to Gs.
This phenomenon was first observed in labs in the 1990s, but for a long time it seemed that the structure could only occur under acidic conditions that did not exist inside a living cell. More recent work has shown the knots could also occur in other environments. On a hunch, Garvan Institute researchers developed an antibody that could sniff out i-motifs in the genome and identify them, tagging them with an immunofluorescent glow.
This allowed researchers to see how frequently and where these knots of DNA occur. They found that the i-motifs are could fold and unfold depending on the acidity of their surroundings, and that the codes were generally found in areas of the genome involved in whether or not a certain gene gets expressed. This suggests the i-motifs may be some kind of switch that can regulate gene expression.
“What excited us most is that we could see the green spots – the I-motifs – appearing and disappearing over time,” says Mahdi Zeraati, the first author of the paper. “It seems likely that they are there to help switch genes on or off.”
Rather shockingly, the I-motif doesn’t just disobey the structural rules, it also disobeys the normally strict base-pairing rules for DNA, which hold that adenine always binds to thymine, and cytosine always hooks up with guanine. In this instance, the structure is formed by two cytosines pairing up.
Given that 98% of the genome does not code for proteins, yet much of this DNA code predicts health and disease, the push is on to decode this information. “Its probably not one dimensional but three or four dimensional”, says Mattick. As Dinger puts it, “These findings will set the stage for a whole new push to understand what this new DNA shape is really for.”
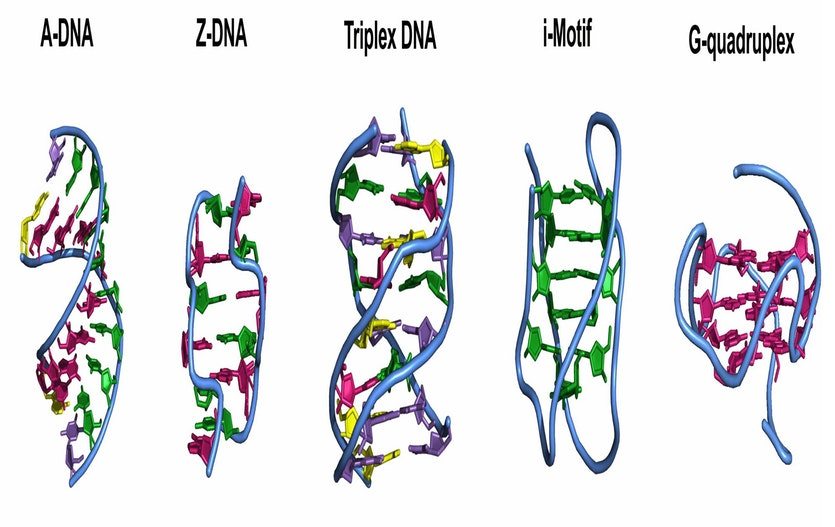
In the future, Zeraati and colleagues will attempt to learn what exact functions these DNA structures serve inside our bodies. Also, perhaps other alternative DNA conformations exist in human live cells, as well, such as A-DNA, Z-DNA, triplex DNA, and cruciform DNA. Such discoveries might open the gates for a new age of genetic research.