CSIR NET Unit 13 Notes – Methods in Biology
Subunit – 13A – Gene Cloning, rDNA Technology
The Last Unit of CSIR NET Syllabus, UNIT 13 – Methods in Biology, deals with different techniques and their applications. Mostly experiment-based questions are asked from this unit that tests your research abilities. Some direct questions are always asked from this unit, especially from subunit D-H. There is too much to cover in this UNIT 13 of CSIR NET life science syllabus, so we suggest you start preparation early, cover important topics, revise as much as possible & don’t forget Biotecnika is there to assist you. Below you can download CSIR NET Unit 13 Notes PDF.
REFERENCE BOOKS For CSIR NET UNIT 13
- Principles of Gene Manipulation and Genomics by Sandy B. Primrose, Richard Twyman
- Gene Cloning and DNA Analysis: An Introduction by T. A. Brown
- Molecular Cloning: A Laboratory Manual by Joseph Sambrook, David Russell
- Fundamentals of Biostatistics by V.B. Rastogi
- Principles and Techniques of Biochemistry and Molecular Biology by Wilson/Walker
RECOMBINANT DNA TECHNOLOGY
- “The joining of DNA molecules from different organisms and inserting it into a host organism to produce new genetic combinations that are of value to science, medicine, agriculture, and industry.”
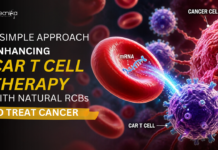
- A clone is a collection of molecules or cells, all identical to an original molecule or cell.
- DNA cloning produces many identical DNA molecules from single ancestral DNA molecules.
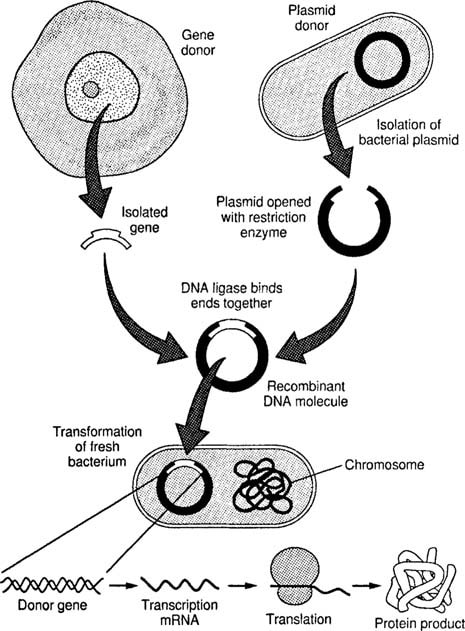
Applications Of Recombinant DNA Technology
- It is used in medicine and research to produce recombinant insulin, growth hormones, hepatitis B vaccines, etc.
- It is used in agriculture to produce golden rice and insecticide-resistant crops.
EXTRACTION OF NUCLEIC ACIDS
Source of DNA
- Distinct kinds of DNA.
- First, total cell DNA is often required as a source of the material (culture of bacteria, from a plant, from animal cells).
- The second type of DNA that is required is pure plasmid DNA.
Download Full Notes For CSIR NET UNIT 13A Notes – Gene Cloning, rDNA Technology
Isolation And Purification of plasmid DNA
Separation based on size
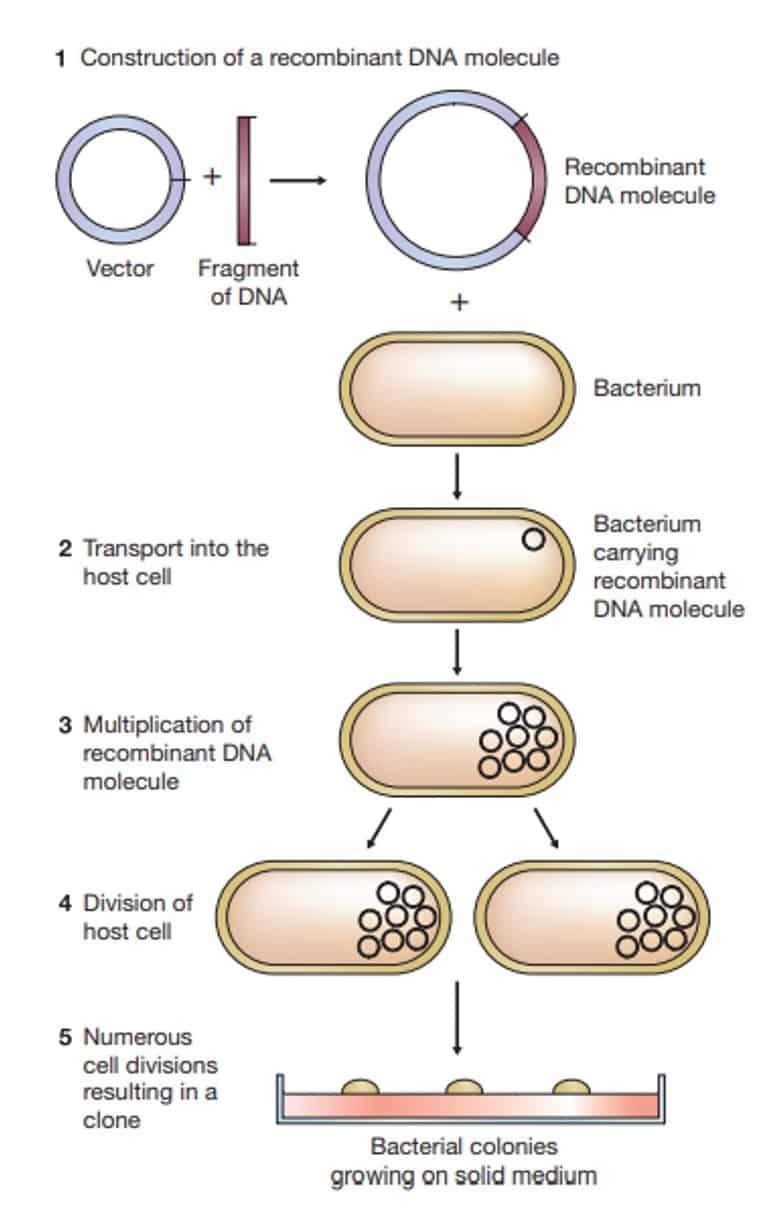
- The usual stage at which size fractionation is performed during the cell extract preparation.
- Only a minimal amount of chromosomal DNA breakage occurs if the cells are lysed under very carefully controlled conditions.
- The resulting DNA fragments are still very large-much, much larger than the plasmids, and can be removed with the cell debris by centrifugation.
- This process is aided by the bacterial chromosome physically attaching to the cell envelope.
2. Separation based on conformation
- Plasmids are supercoiled molecules formed by the partial unwinding of the double helix of the plasmid DNA during the plasmid replication process by enzymes called topoisomerases.
- The supercoiled conformation can be maintained when both polynucleotide strands are intact, called covalently closed-circular (ccc) DNA.
- If one of the polynucleotide strands is broken, the double helix reverts to its normal relaxed state, taking an alternative conformation called open-circular (oc).
- Super coiling is important in plasmid preparation due to the easy separation of supercoiled molecules from non-supercoiled ones.
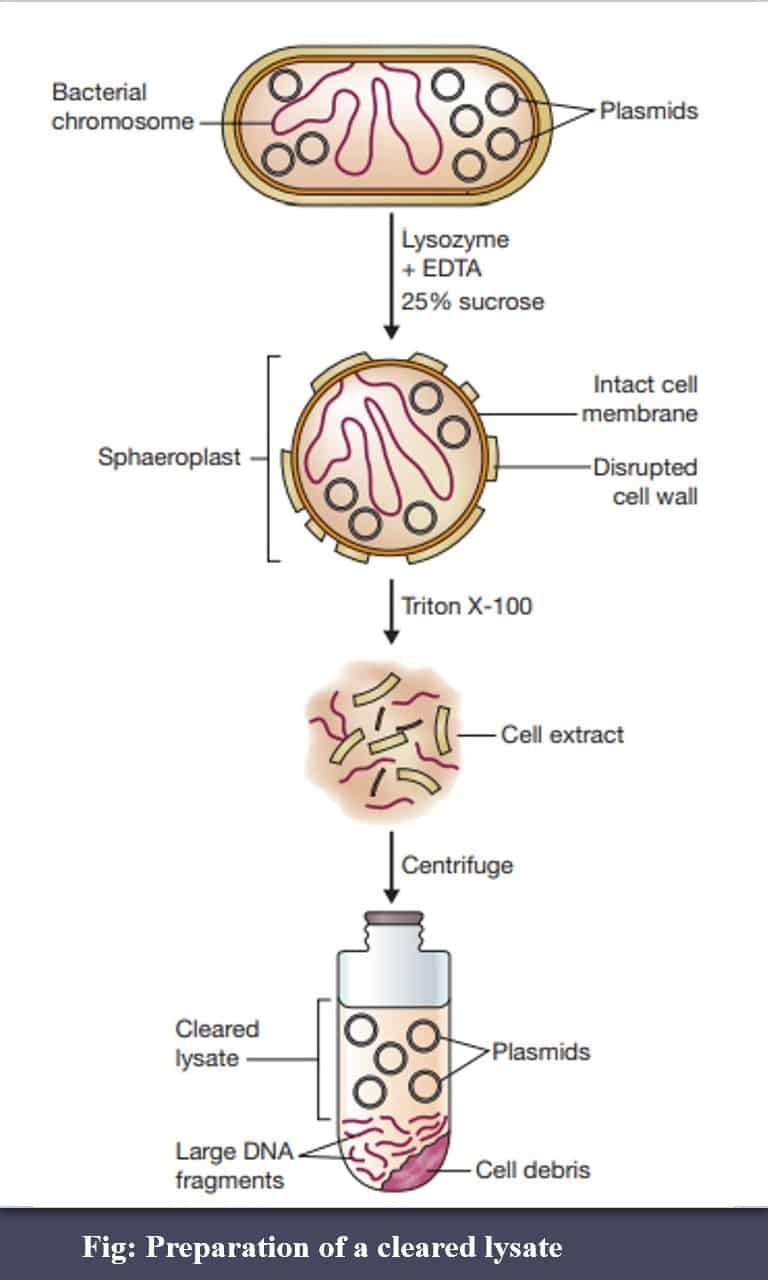
- Two conformations of circular double-stranded DNA:
- supercoiled—both strands are intact;
- open-circular —one or both strands are nicked.
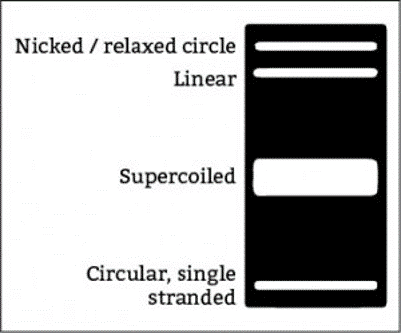
These are the three forms of the plasmid:
- Supercoiled (smallest 3D structure, runs fastest.
- Linear (runs in the middle) and relaxed, nicked plasmid (runs highest).
- Another fourth band, which is the circular single-stranded plasmid (runs faster than the supercoiled version)
Alkaline denaturation method
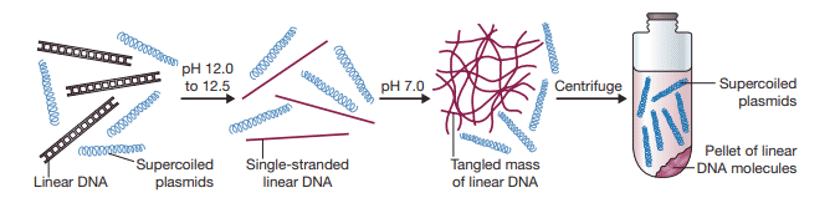
- This method is based on maintaining a very narrow pH range for denaturing non-supercoiled DNA but not the supercoiled plasmid.
- Adding sodium hydroxide to cell extract or cleared lysate (pH 12.0-12.5) disrupts the hydrogen bonds of non-supercoiled DNA molecules.
- As a result, the double helix unwinds, and two polynucleotide chains separate.
- Further addition of acid causes the aggregation of these denatured bacterial DNA strands into a tangled mass which can be pelleted by centrifugation, leaving plasmid DNA in the supernatant.
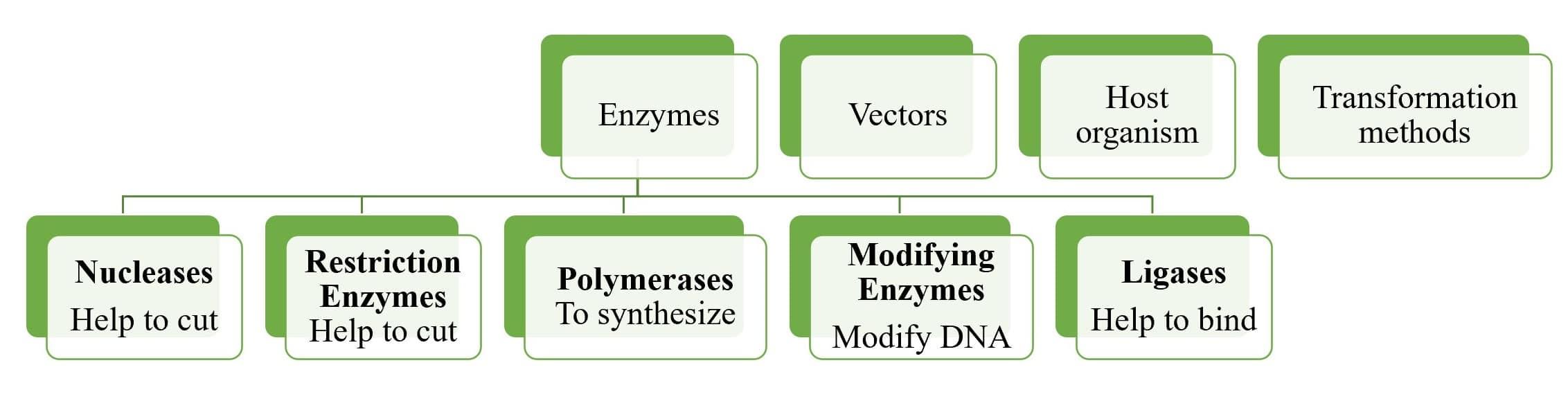
TOOLS OF RDT
DNA Manipulating Enzymes
- Nucleases
- Nucleases degrade DNA molecules by breaking the phosphodiester bonds that link one nucleotide to the next in a DNA strand.
- There are two different kinds of nucleases:–Exonucleases remove nucleotides one at a time from the end of a DNA molecule.
–Endonucleases can break internal phosphodiester bonds within a DNA molecule. - When a double-stranded molecule is attacked, the main distinction between different exonucleases lies in the number of degraded strands.
Liking the notes so far? For notes, PPTs, PDFS on vectors for plants – Ri plasmid, Genomic & cDNA Library, Flowcharts & PPTs for all of the above topics – Join Biotecnika Today. You can opt for your study materials, Online portal, Classroom Coaching, and Online coaching and pave your path towards qualifying for CSIR NET this year with a good rank. Call Toll-FREE – 1800-1200-1818 For More details or chat with us here.
DIFFERENT KINDS OF NUCLEASE
Exonuclease
- Bal31, ((purified from Alteromonas espejiana) removes nucleotides from both strands of a double-stranded molecule. Contd…
Download Full Notes For CSIR NET UNIT 13A Notes – Gene Cloning, rDNA Technology
Or
CSIR NET Life Science PART A Formula Sheet Download + Preparation Strategy
Download Full Notes For CSIR NET UNIT 13A Notes – Gene Cloning, rDNA Technology. CSIR NET UNIT 13 Notes PDF.