CSIR NET DNA Replication Notes
Unit 3 – Molecular Biology: The most common choice of almost all students. If you want to start your CSIR NET preparation then this unit is perfect to start with. Questions are asked from this UNIT almost every time in the CSIR NET Exam and are to the point & direct – subject-specific questions. Here we will be discussing UNIT 3(A) – DNA Replication.
Reference Book For UNIT 3A: DNA Replication
• Lehninger’s Principles of Biochemistry by Nelson & Cox
• Biochemistry by Voet & Voet
• Molecular Biology of the Cell by Alberts
• Molecular Biology by Robert F. Weaver
• Molecular Biology of the Gene by Watson
Four requirements for DNA to be genetic material
- Must carry information – Cracking the genetic code
- Must replicate – DNA replication
- Must allow for information to change – Mutation
- Must govern the expression of the phenotype – Gene function
Mechanisms by which information is transferred in the cell are based on “Central Dogma”
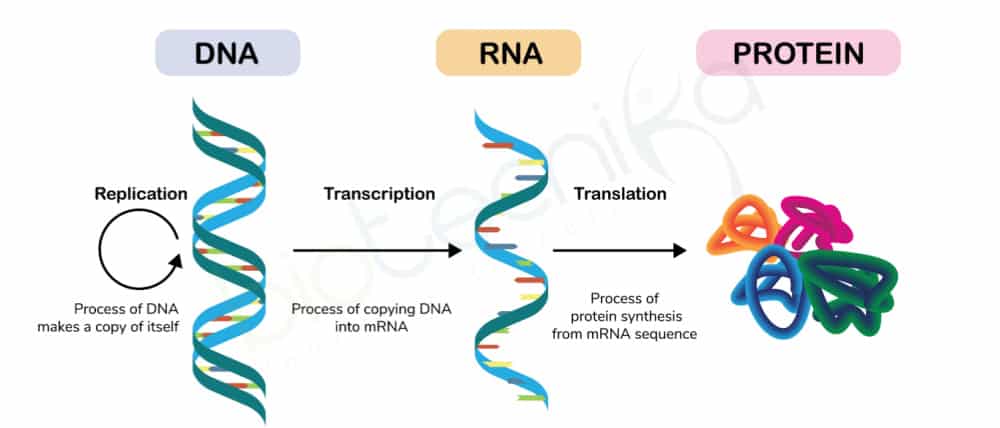
- Replication: synthesis of daughter DNA from parental DNA
- Transcription: synthesis of RNA using DNA as the template
- Translation: protein synthesis using mRNA molecules as the template
- Reverse transcription: synthesis of DNA using RNA as the template
Rules of DNA Replication
- Takes place in the S phase of the cell cycle
- Occur once per cell cycle
- Starts from the origin of Replication
- DNA Replication takes place from 5’to 3’direction
- DNA Replication can be uni or bidirectional
DNA replication 3 possible models
Semi-conservative replication
One strand of duplex passed on unchanged to each of the daughter cells. This ‘conserved’ strand acts as a template for the synthesis of a new, complementary strand by the enzyme DNA polymerase.
Download Full PDF – CSIR NET UNIT 3A Notes – DNA Replication
How do we know that DNA replication is semiconservative?
The Meselson-Stahl Experiment
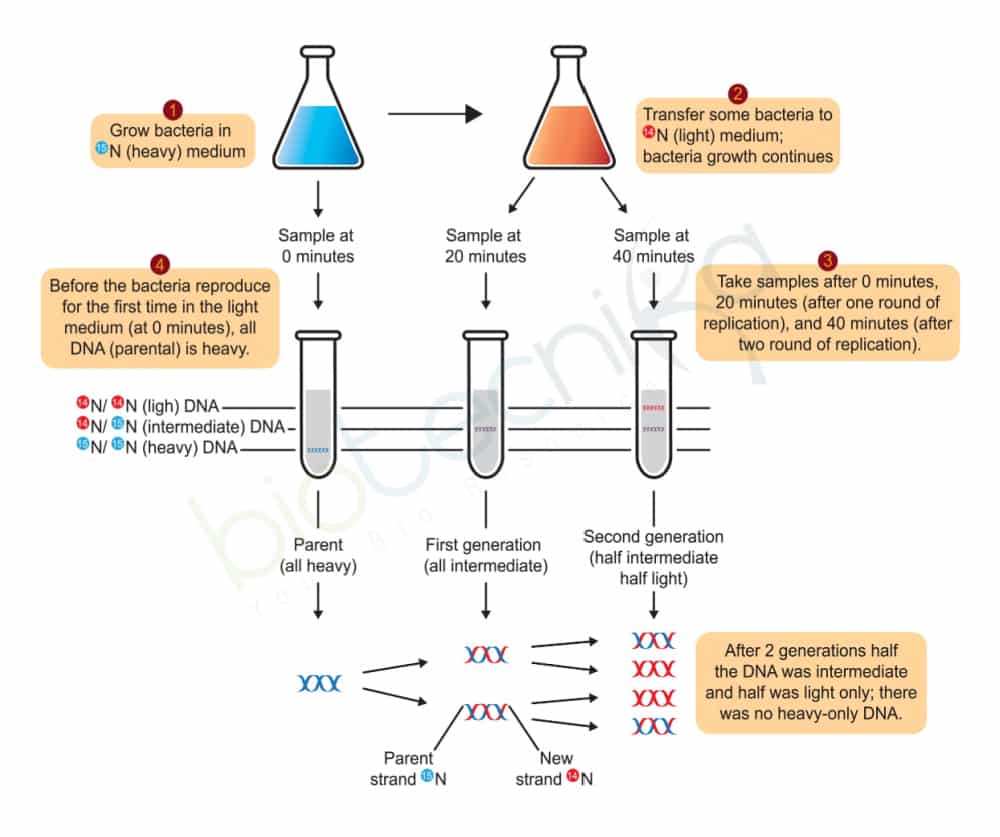
- Meselson and Stahl (1958) grew E. coli in a heavy (not radioactive) isotope of nitrogen, 15N in the form of 15NH4Cl. Because it is heavier, DNA containing 15N is denser than DNA with normal 14N, and so can be separated by CsCl density gradient centrifugation.
- Once the E. coli were labeled with heavy 15N, the researchers shifted the cells to a medium containing normal 14N and took samples at time points. DNA was extracted from each sample and analyzed in CsCl density gradients (Figure 3.2).
Uni or bidirectional
- Replication starts from unwinding the dsDNA at a particular point (called origin),
followed by the synthesis of each strand.
- The parental dsDNA and two newly formed dsDNA form a Y-shape structure called a replication fork.
Bidirectional replication
- Once the dsDNA is opened at the origin, two replication forks are formed spontaneously.
- These two replication forks move in opposite directions as the syntheses continue.
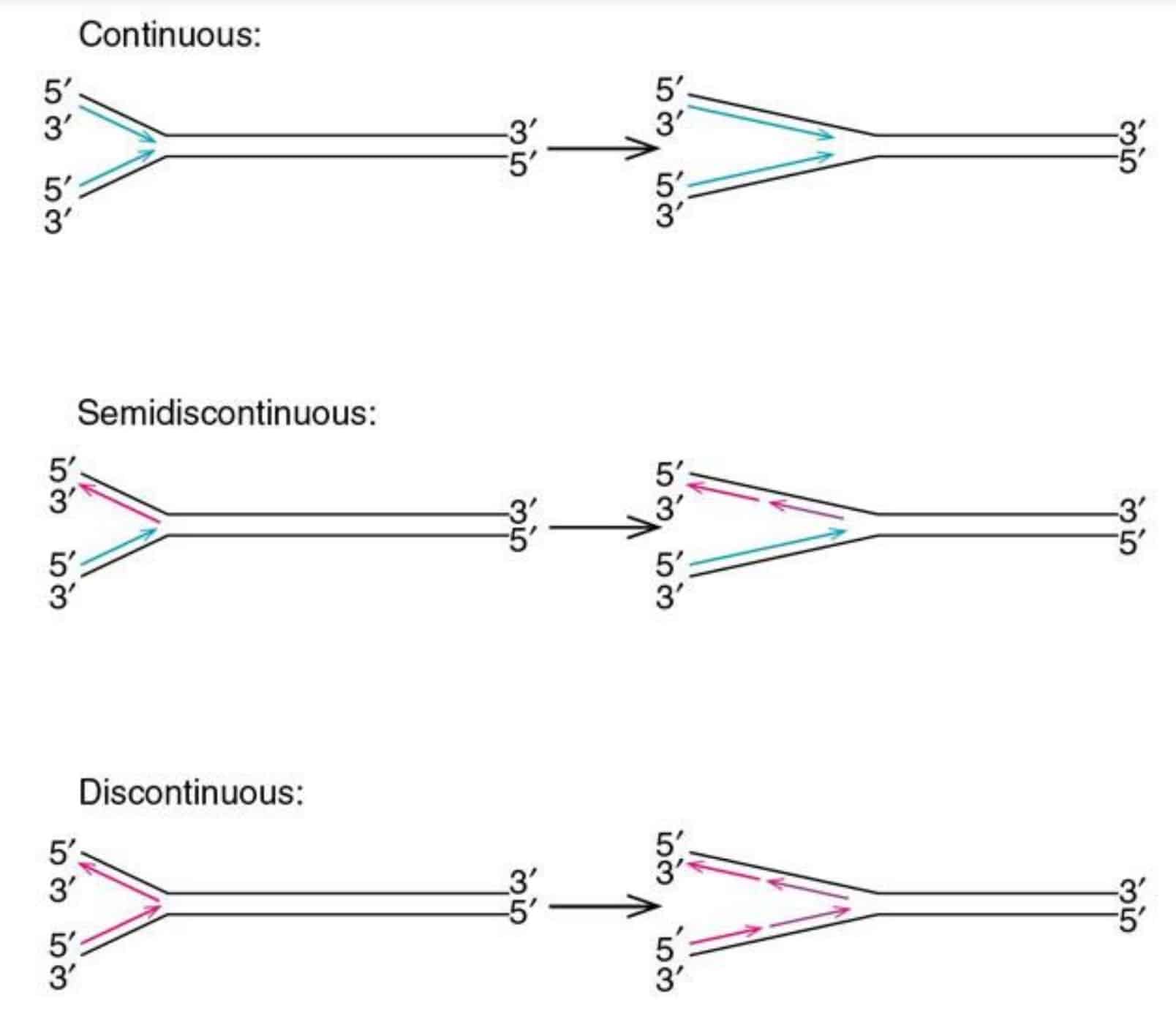
Semi-continuous Replication
The daughter strands on two template strands are synthesized differently since the replication process obeys the principle that DNA is synthesized from the 5´ end to the 3´end.
New strand synthesis is always in the 5’-3’ direction.
Discovery of Okazaki fragments
Evidence for semi-discontinuous replication [3H] thymidine pulse-chase labeling 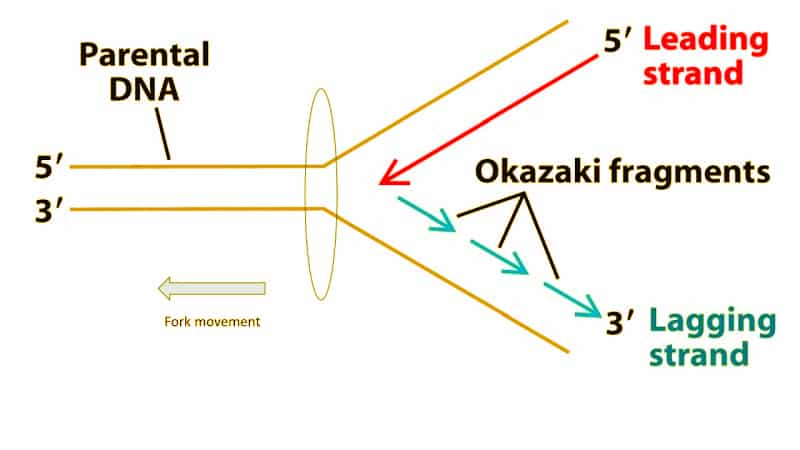
1. Grow E.coli
2. Add [3H] thymidine in the medium for a few seconds spin down and break the cell to stop labeling analyze found a large fraction of nascent DNA (1000-2000 nt) = Okazaki
fragments
3. Grow the cell in a regular medium then analyze the small fragments join high molecular weight DNA= Ligation of the Okazaki fragments
Enzymes required for DNA replication
More than 20 different enzymes are involved in DNA replication. The entire complex is
termed as REPLISOME….Contd.
Detailed Notes on DNA Replication including Eukaryotic DNA Replication – Available which includes:
- Flow charts
- Koncept Tables
- Lecture Videos
- Question Discussion
- Doubt Solving
Join Josh Batch December 2022 Starting on 6th June
Chat with us to Book your seat Today
Get a FREE Computational Biology Virtual Internship opportunity,
JCB Scholarships for Talented candidates,
FREE Koncept Notes, and Much more…
Download Full PDF – CSIR NET UNIT 3A Notes – DNA Replication
Download CSIR NET Notes PDF
- UNIT 1(A) Notes – Structure of Atoms, Molecules & Chemical Bonds – https://btnk.org/unit1a
- UNIT 1(B) Composition, Structure & Function of Biomolecules – https://btnk.org/unit1b
- UNIT 1(G) Ramachandran Plot – https://btnk.org/unit1g
- UNIT 2 Organization of Genes & Chromosomes – https://btnk.org/unit2
- UNIT 2D Cell Division & Cell Cycle – https://btnk.org/unit2d





















 followed by the synthesis of each strand.
followed by the synthesis of each strand.














