Animals At Risk Of Coronavirus Infection Identified Using Genome Analysis
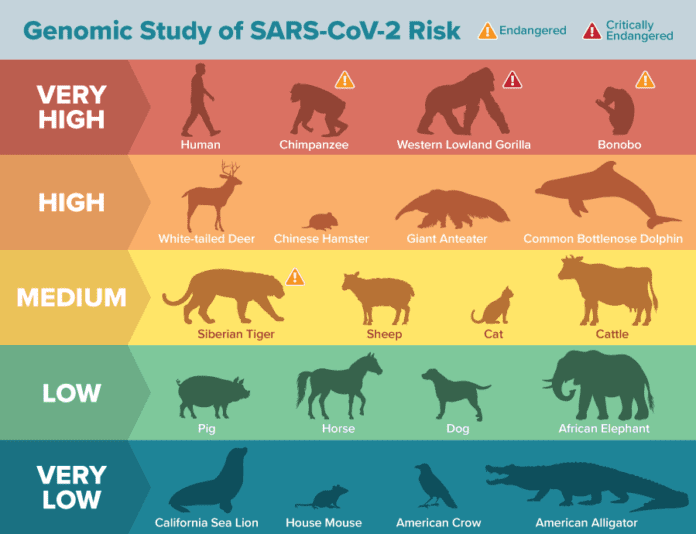
According to a study conducted by an international team of scientists at the University of California, Davis (UC Davis), humans are not the only species facing a potential threat from SARS-CoV-2. The team compared the amino acids of a key part of the ACE2 protein that is used by SARS-CoV-2 to gain entry into human cells with that of more than 400 animals using genome analysis and found that different species, including the highly endangered Western lowland gorilla, could potentially be infected with the novel coronavirus.
The study could help to identify intermediate hosts for SARS-CoV-2, even though they caution not to overinterpret their results, which are based on computational models. Harris Lewin, Ph.D., lead author for the study, said the data offers a starting point to identify threatened and vulnerable animal populations at risk of COVID-19 and could inspire practices that protect both animal and human health. The results were published in the Proceedings of the National Academy of Sciences (PNAS).
Previously, comparative genome analysis had shown that SARS-CoV-2 probably had ancestors that originated in bats and then could have jumped to an intermediate host from where viruses extended host range to mammals and primates. Coronaviruses cause infections in many mammalian species and are associated with severe clinical diseases like enteric and respiratory diseases in pigs and cattle.
The researchers suggested that the pandemic can be better predicted and controlled with the understanding of the host range of SARS-CoV-2.
Angiotensin I converting enzyme 2 (ACE2) is the receptor for SARS-CoV-2 spike (S) protein in humans. ACE2 is found in several types of tissues and cells, including epithelial cells in the mouse, nose, and lungs. In the ACE2 protein, 25 amino acids are crucial for the virus to bind to cells and invade them. To study how many of these amino acids are found in the ACE2 protein of the other species, the researchers used these 25 amino acid sequences of the ACE2 protein, their predicted protein structure, together with the coronavirus spike protein.
They used a combination of protein structural analysis and comparative genomic approaches to study the potential of ACE2 homologs from 410 vertebrate species (including species from amphibians, reptiles, fishes, and mammals) to serve as a receptor for the novel coronavirus.
The International Union for Conservation of Nature classed 40% of the potentially susceptible animals as “threatened” and may be vulnerable to human-to-animal transmission. These include many critically endangered primate species like the Sumatran orangutan, Northern white-cheeked gibbon, and Western lowland gorilla. Marine animals like bottlenose dolphins, grey whales, and Chinese hamsters are also at high risk of coronavirus infection.
In the known cases of coronavirus infection in cats, mink, dogs, lions, hamsters, and tigers, the virus may be using ACE2 receptors or any other receptor to invade the host cells. The lower propensity for binding could lead to the lower ability of the infection to spread or lower propensity for coronavirus infection. Institutions, including the San Diego Zoo and the National Zoo, contributed genomic material to the study and strengthened programs to protect both animals and humans.
Further study is required to understand how the findings from the study relate to disease or infection risk, but the correlation is high for those species that have known infectivity data.
Co-author Klaus-Peter Koepfli, Ph.D., a senior research scientist at Smithsonian-Mason School of Conservation, said the new information has allowed them to focus their efforts and plan accordingly to keep humans and animals safe.
However, the researchers also cautioned against overinterpretation of the predictions solely based on in silico analyses, saying additional experimental data is required to confirm actual risks. As more extensive data are generated, the prediction accuracy of the model may be improved.
Research had shown that the immediate ancestor of the novel coronavirus originated in bats. But bats are at low risk of contracting the virus through the ACE2 receptor, consistent with the actual experimental data. Whether bats transmitted the virus to humans through an intermediate host or directly is not revealed yet, but the new findings support the idea of intermediate hosts.
The research concluded by saying their data could help to identify the intermediate species of the coronavirus and protect both animals and humans from the virus.