Molecular Interaction Study For New Medicines
Researchers at the University of Minnesota have carried out a first-of-a-kind study on molecular interaction. This research will make it easier and more efficient for scientists to develop new medicines and other therapies for diseases such as HIV AIDS, cancer, and autoimmune diseases.
The research resulted in a mathematical framework that simulates the effects of the critical parameters that control interactions between molecules that have multiple binding sites, as is the case for many medicines. Scientists at the University of Minnesota plan to use this computational model to develop a web-based application that other scientists can use to escalate the development of new therapies for diseases.
Molecular Interaction Study For New Medicines- The New Methodology
Casim Sarkar, a University of Minnesota biomedical engineering associate professor as well as the senior author of the study, said that this is the first time a mathematical model has been used for this kind of research instead of the conventional trial and error experimental method.
This computational model will make research highly efficient & could accelerate the creation of new therapies for many kinds of diseases.
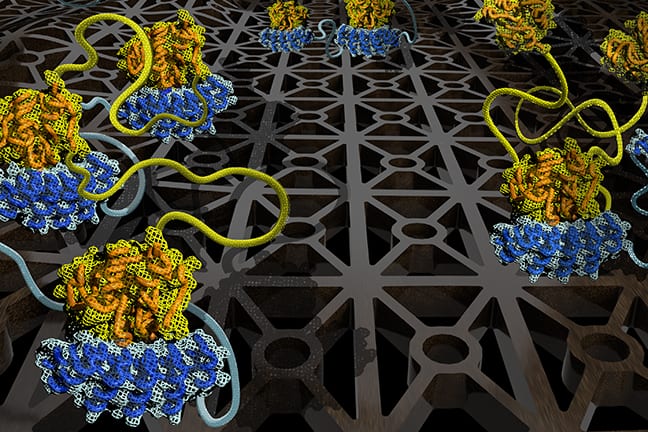
The team of scientists studied three main parameters of molecular interactions—binding strength of each site, the rigidity of the linkages between the sites, as well as the size of the linkage arrays.
The scientists looked at how these three parameters can be “dialed up” or “dialed down” to control how molecule chains with two or three binding sites interact with one another. The research team then confirmed their model predictions in lab experiments.
Molecular Interaction Study For New Medicines- Maths For Medicine
The researchers highlight the need for a mathematical framework to decode this programming language. The study notes that even when the interacting molecule chains have just three binding sites each, there are a total of 78 unique binding configurations. These binding configurations cannot be experimentally observed.
By dialing the parameters in this new mathematical model, scientists can quickly understand how these different binding configurations are affected & tune them for a wide range of biological and medical applications.
Wesley Errington, a University of Minnesota biomedical engineering postdoctoral researcher & lead author of the study, said that this new mathematical model could be applied to more complex molecule interaction.