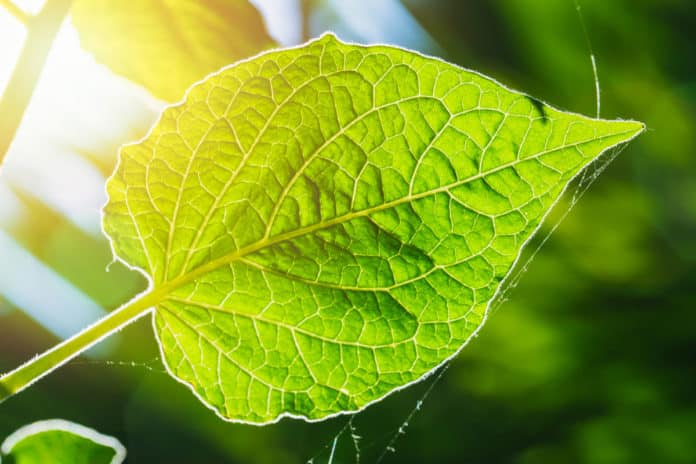
21 New Genes Discovered That Regulate Photosynthesis
Princeton University researchers by utilizing a public library they created to help the research community search & identify genes and their functions – have identified 303 genes related to photosynthesis including 21 newly discovered genes with high capability to provide fresh insights into the life-sustaining biological process. The research was published this week in Nature Genetics.
“The region of the plant in charge of photosynthesis is similar to an intricate machine made up of several components, and we want to understand what each part does,” said Martin Jonikas, assistant professor of molecular chemistry at Princeton. “This library we expect will be one of the foundations that people will assemble on to make the next generation of discoveries”
Unlocking the role of each gene could allow investigators to engineer plants to grow more quickly, possibly meeting future world food needs. Plants may also possibly be altered to absorb more carbon dioxide, helping to deal with climate challenges.
Each “book” in the library is really a breed of Chlamy using a single mutation.
Libraries of a similar kind have been developed in other unicellular organisms like yeast, but this approach remains one of its kind for any single-celled photosynthetic organism. Unicellular organisms because of their rapid growth capability are excellent research tools.
“Because this algal species is frequently utilized as a model to understand a wider array of biological processes, this library is going to be a significant resource,” said Karen Cone, a program manager in the National Science Foundation, which was the key funder for this research.
The project took two long years to finish. Throughout the project, researchers used robots to maintain generations of cells living by altering the nutrient-rich liquid media where the cells live.
Back in 2010, the project was initiated. Jonas and his team were in Carnegie Institution for Science on the Stanford University campus and was completed at Princeton where the Jonikas laboratory moved in 2016.
The project was run in coordination with a senior staff scientist at Carnegie – Arthur Grossman & with the Chlamydomonas Resource Center conducted by Paul Lefebvre, a professor of plant and microbial biology at the University of Minnesota.
The capacity to watch a Chlamy mobile with only one defective gene among all the other working genes allows researchers to determine what that gene does. As an instance, if the mobile has trouble transferring, then the faulty gene’s function mostly probably entails regulating motion.
Jonikas compared the Chlamy mutant library to a library containing many copies of a guide for constructing a vehicle, with each copy of this book missing a different section. No matter which manual was used, the subsequent car would be missing a part, making it unable to function as anticipated.
“The horn might not work, or the steering wheel may not turn,” Jonikas stated. “Then you would know that the missing segment included the directions for that part of the automobile.”
The library enables investigators to test multiple mutant Chlamy breeds at once because each mutation is tagged with a distinctive “DNA barcode.” For the current study, investigators placed tens of thousands of Chlamy strains in a single flask and exposed them to light. Strains that failed to grow were prone to contain a gene involved in photosynthesis.
One of the recently identified genes is CPL3, which is considered to play a role in amassing the protein “components” of the photosynthetic machinery. The group is now exploring whether the gene aids the algae to adapt their photosynthetic activity to changes in sunlight amounts.
The mutant library may empower studies in different areas of plant biology, such as intracellular communication and Chlamy’s capability to paddle about its surroundings using a tail-like cilium.
Xiaobo Li, the study’s first author, was a postdoctoral researcher in Princeton when the team finished the library. “It is our expectation that the Chlamydomonas mutant library along with the genes identified will result in numerous basic discoveries in photosynthesis, cell motility and lots of different procedures,” Li explained.
Weronika Patena, a senior bioinformatics analyst at the Jonikas laboratory, wrote computer programs to examine large amounts of information to identify genes likely to be involved in photosynthesis. “I believe the success of the project will greatly accelerate research into photosynthesis and other processes for which Chlamy is a good model and supply a great deal of value to the scientific community,” she said. This project was supported by a grant from the National Science Foundation (MCB-1146621), the National Institutes of Health (DP2-GM-119137), the Simons Foundation and the Howard Hughes Medical Institute (55108535), the European Molecular Biology Organization (ALTF 1450-2014 and ALTF 563-2013), the Swiss National Science Foundation, along with Westlake University in China.