Impaired “Epigenetic Landscape” Hallmark of Alzheimer’s : Study
Alzheimer’s Disease (AD) is the most common cause of dementia in the elderly. A complex interaction between genetic and environmental factors likely contributes to the molecular processes that drive AD.
Although genetic variation in specific genes increases the risk of AD, age is the strongest known risk factor. How molecular processes of aging predispose to AD, or become deregulated in AD, remains to be understood.
It is seen in animal models, that epigenetic marks such as histone acetylation are associated with learning and age-related memory decline. Histone acetylation is reduced at memory genes in mouse models for AD, and treatments with nonselective histone deacetylase inhibitors aiming to reverse loss of acetylation have shown promising results in restoring synaptic and cognitive plasticity in mouse models of AD.
Among the histone acetylation marks, H4K16ac is a key modification because it regulates chromatin compaction, gene expression, stress responses and DNA damage repair. In model organisms, modulators of H4K16ac play a role in whole organism aging and cellular senescence. Also, senescent cells display H4K16ac enrichment over promoter regions of expressed genes.
Therefore, the scientists at the Perelman School of Medicine at the University of Pennsylvania hypothesized that epigenetic regulation by H4K16ac may be involved in aging of the human brain and perhaps in the progression of AD.
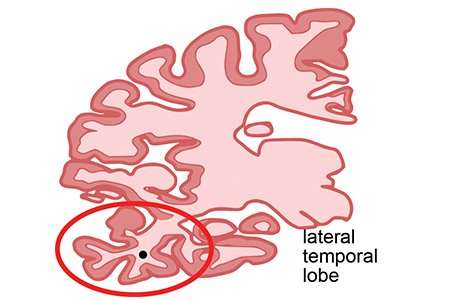
They therefore profiled the epigenomic landscape of AD brains, specifically in one of the regions affected early in AD, the lateral temporal lobe. They compared these to both younger and elderly cognitively normal control subjects. The team described the genome-wide enrichment of a chemical modification of histone proteins that regulates the compaction of chromosomes in the nucleus (called acetylation of lysine 16 on histone H4, or H4K16ac for short).
“This is the first time that we have been able to look at these relationships in human tissue by using donated postmortem brain tissue from the Penn Brain Bank,” said Shelley Berger, PhD, a professor of Cell and Developmental Biology in the Perelman School of Medicine and a professor of Biology in the School of Arts and Sciences. “Our results establish the basis for an epigenetic link between aging and Alzheimer’s disease.”
A three-way comparison of younger, older, and AD brain tissue revealed a specific class of H4K16ac changes in AD compared to normal age-established changes in the brain. This finding indicates that certain normal aging changes in the epigenome may actually protect against AD and when these goes awry, a person may become predisposed to AD.
“These analyses point to a new model of Alzheimer’s disease. Specifically it appears that AD is not simply an advanced state of normal aging, but rather dysregulated aging that may induce disease-specific changes to the structure of chromatin – the combination of histone proteins and DNA.” said first author Raffaella Nativio, PhD, a postdoctoral fellow in Berger’s lab.
The team next hopes to discover the physiological changes that cause the decrease of H4K16ac specifically in AD brains, but not in normal-aged brains.