Flu Genome Could Hold Clues for Better Pandemic Flu Surveillance
Generally very slow, evolution in case of viruses remains fast and deadly. Flu viruses change rapidly to escape the body’s defenses. Every few years, new variants of flu emerge and cause epidemics around the world.
In the past several years, advances in genome sequencing have begun to shed light on the beginnings of viral evolution, deep within individual infections. And scientists at the Washington University School of Medicine in St. Louis wondered whether, for flu, this information might give us an early glimpse of future global evolutionary trends.
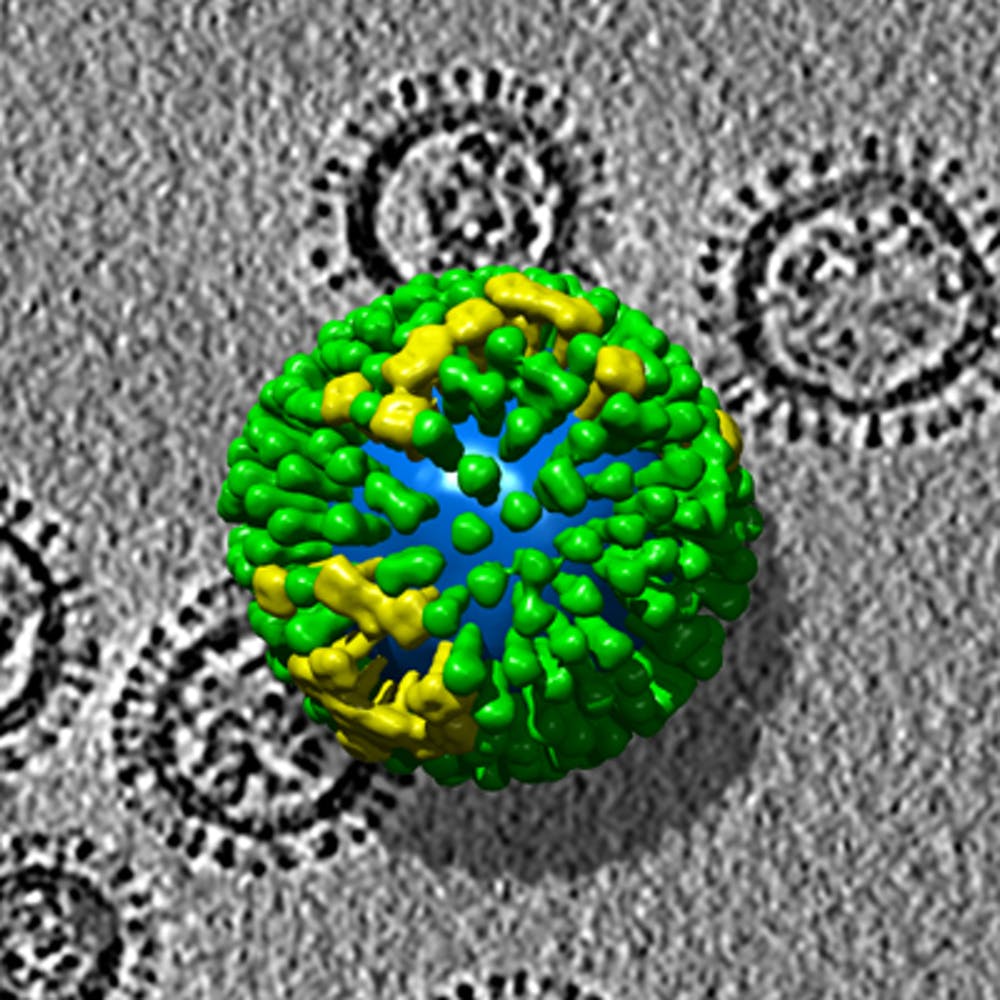
Influenza A virus nucleoprotein (NP) association with viral RNA (vRNA) is essential for packaging, but the pattern of NP binding to vRNA is unclear. Influenza A virus (IAV) possesses a segmented, negative-sense RNA genome that is bound by the viral nucleoprotein (NP) throughout replication. However, current models conflict with each other and yield no information about RNA conformation, binding, or NP–RNA association.
Therefore, the scientists in the study accessed the native-state of NP–vRNA interactions in infected human cells in order to assess if a genetic mix of strains could produce a hybrid virus with the potential to explode into pandemic flu.
“We think that two strains need to have similar features in their genome to re-assort and make a new virus,” said senior author Jacco Boon, PhD, an assistant professor of medicine at Washington University. “We hope that in the future, this work will allow us to focus on certain strains of influenza virus and target our surveillance more narrowly so we have a better chance of identifying the next pandemic flu before it spreads.”
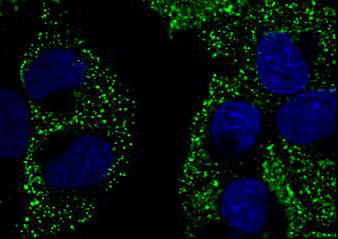
The scientists began with Madin-Darby Canine Kidney cells, a line that is uniquely tailored to be a mammalian model for studying virus infections. The cells were then infected with both contemporary strains of Influenza A – such as a strain of H1N1, and H7N3 and some other mutants – with historical viral samples.
The mixing and matching of the DNA was then assessed by the PAR-CLIP method of looking at the binding sites, in conjunction with next-generation sequencing.
The flu genome is broken into eight parcels of RNA, they demonstrated. Those parcels are contorted into 3-D shapes of genetic information. When the mutations change the shape, it compromises the viral replication. The result was a new influenza strain born with genetic information from multiple parental strains.
“Right now we do surveillance on pretty much everything,” said Boon, who is also an assistant professor of molecular microbiology, and of pathology and immunology. “But if we know that the viruses from a certain species or a certain region just don’t have the right RNA features, then we can make surveilling them a lower priority. If we can focus our resources more effectively, we may be able to catch the next pandemic flu before it really gets going.”