Analysis of Pixelated Spatial Gene Expression Allows for Rapid Cancer Identification
The spatial localization of gene expression can unravel important insights into tissue heterogeneity, functionality, and pathological transformations, but the ability to maintain this spatial information remains an enduring challenge in tissue sections routinely used for pathology. Amplification-based spatial gene expression analysis methods provide good sensitivity and specificity but decouple the analyte isolation and biochemical detection steps, making them low throughput and laborious.
Direct probe-based hybridization techniques such as single-molecule fluorescence in situ hybridization (FISH) allow direct visualization of single RNA molecules in their native cellular context but are not amenable to performing this on tissue section in a high throughput manner.
Methods to perform spatially mapped transcriptome analysis on a tissue section can identify multiple targets simultaneously but they must trade-off between the histologic reference and the quality of recovered biomaterials, as staining and manual identification are often needed
These constraints limit the translation of the above methods into routine research and clinical practice.
Therefore, in this direction a University of Illinois and Mayo collaboration has demonstrated a novel gene expression analysis technique that can accurately measure levels of RNA quickly and directly from a cancerous tissue sample while preserving the spatial information across the tissue.
According to Illinois Bioengineering Professor Rashid Bashir, existing gene expression methods have limitations. “They are cumbersome and slow, taking hours or even many days to do the analysis across just one tissue sample,” said Bashir, co-leader of the research team, who is a Grainger Distinguished Professor of Bioengineering and the Carle Illinois College of Medicine executive associate dean. “Our technique does the entire analysis across the tissue slice in two hours or less.”

The technique improves upon the of conventional methods by analyzing a starting sample of tissue cryosection and performing parallel picoliter reverse transcriptase loop mediated isothermal amplification (RT-LAMP) reactions with minimal sample processing.
Research team co-lead Dr. Farhad Kosari, an assistant professor of biochemistry and molecular biology at Mayo Clinic, envisions their new technique could someday be used in research labs as well as the clinic. “If you were studying tumor microenvironments, you’d want to know what genes are expressed at a specific location of the tumor,” said Kosari. “This ability to look at localized expression of genes is currently done with fluorescence in situ hybridization (FISH), but our technique is a lot faster and more quantitative.”
In addition, the messenger RNA (mRNA) will elicit more accurate genetic information. “When analyzing animal models and xenografts, there can be cross reactivity of the target antibody with the host antigen leading to false positive signals and error in analysis,” said Illinois Bioengineering doctoral student Anurup Ganguli, the first author of the study.
In the course of their study, the team devised a fingernail-size silicon chip that contains an array of more than 5,000 pyramid-shaped wells with razor-sharp edges. When a centimeter-sized cancer tissue sample is placed on the chip it is automatically cut up into hundreds or thousands of tiny pieces that are analyzed in parallel using an existing technology known as loop mediated isothermal amplification (LAMP).
LAMP is an isothermal reaction which has been shown to be robust against factors in tissue that inhibit a polymerase chain reaction (PCR). It uses 4–6 primers which identify 6–8 regions on the template for amplification which makes it more specific than PCR.
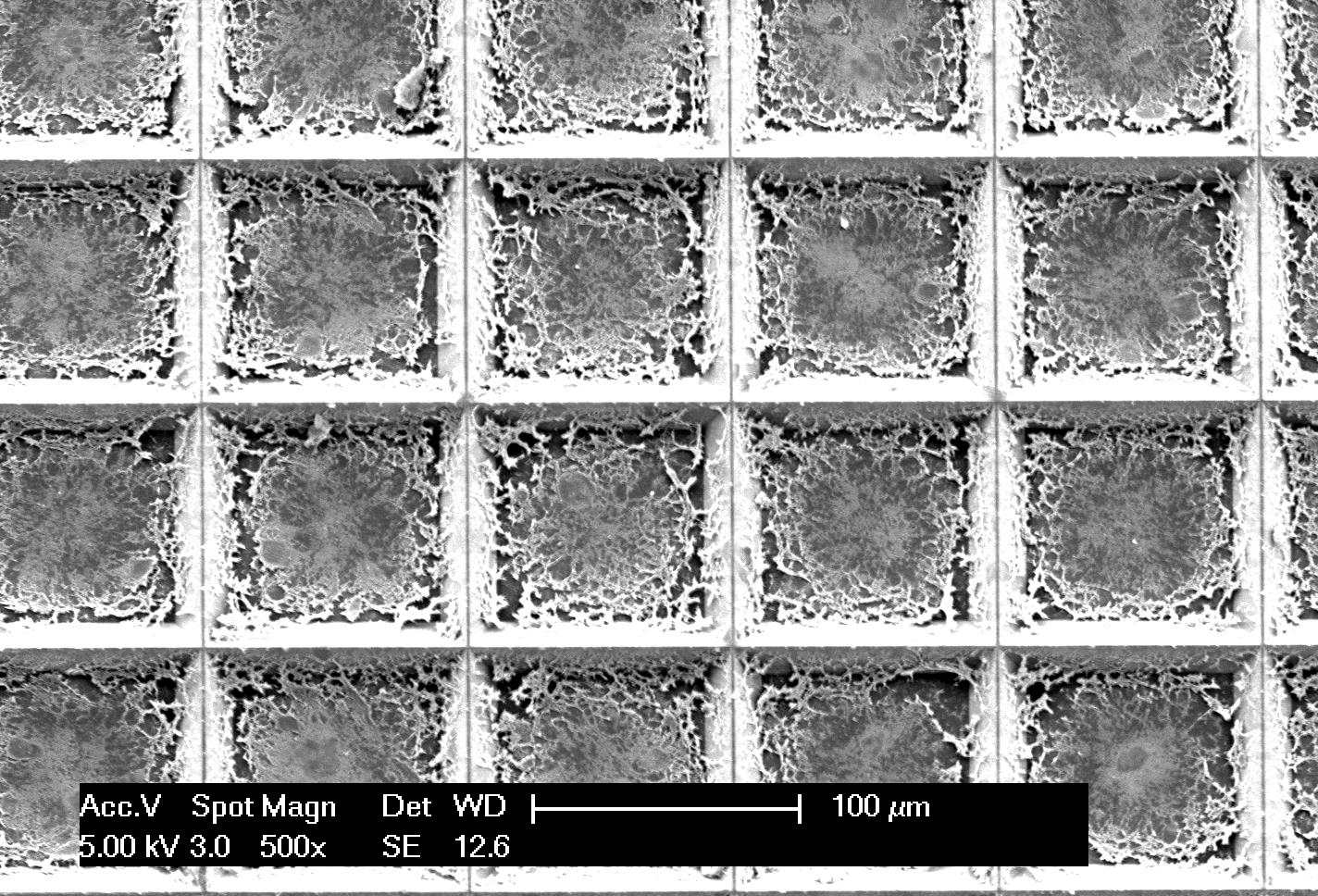

Once the tissue is cut and placed into the underlying wells on the microchip, thousands of independent picoliter volume LAMP reactions are performed in each well directly from the tissue without the need for any analyte purification.
“Laser capture microdissection (LCM) followed by downstream purification and amplification has been used in the past to look at specific regions of stained tissue samples and analyze the heterogeneity within the sample,” Kosari said. “Our technique is similar to performing more than 5,000 LCM steps with the downstream amplification in a single step on a microchip.”
The technique was demonstrated using frozen human prostate tissue xenografts grown in mice.
In less than two hours, the team was able to amplify and analyze the mRNA of TOP2A, a nuclear enzyme and known marker of prostate cancer’s aggressiveness.
“Our approach pixelates the entire tissue sample and can identify those very few cells that may be cancerous,” said Bashir. “There isn’t any technique in existence that takes raw tissue to nucleic acid amplification while keeping the spatial information preserved.”
The team will further work on the gene expression technique and aims to use it to map genetic mutations for lung and breast cancers, in addition to prostate cancer.